Theme 11 — Drug Repurposing and Synthetic Lethality
"The cheapest drug is the one that already exists. The hardest question is knowing when to use it differently."
Corpus snapshot: 6,386 papers · 0 Tier A · 24 Tier B
The Thinnest Evidence Base
Of all the thematic clusters in this corpus, drug repurposing commands the thinnest evidence base relative to its potential clinical impact. With zero Tier A papers and only 24 Tier B publications, this theme occupies the most sparsely populated tier of the landscape — a striking deficit given the practical urgency of expanding the therapeutic arsenal for NSCLC beyond the current menu of targeted agents and immunotherapies. The deficit is not for lack of computational tools: connectivity mapping, network pharmacology, and gene-drug interaction databases have matured considerably over the past two decades. Rather, the bottleneck lies in the translational gap between computational prediction and clinical validation, and in the field's overwhelming preference for de novo drug development over the methodical repurposing of existing pharmacopoeia. This chapter examines what the available evidence supports, identifies the critical transition from traditional drug-matching frameworks (DMF) to synthetic lethality-based approaches, and argues that this theme represents one of the highest-yield opportunities for expansion in computational lung cancer research.
Connectivity Mapping: The Foundational Platform
The intellectual architecture of computational drug repurposing rests substantially on the Connectivity Map (CMap), the seminal platform developed by Lamb and colleagues that enabled researchers to query gene expression signatures against a reference library of drug-induced transcriptomic perturbations [PMID: 17186018]. The logic is elegant in its simplicity: if a disease state produces a characteristic gene expression signature, then a drug whose transcriptomic effect is inversely correlated with that signature may reverse the disease phenotype. The CMap approach has since been extended to encompass tens of thousands of perturbagens across hundreds of cell lines, creating a dense reference matrix for hypothesis generation. Zhao and colleagues applied CMap-based reasoning specifically to oncotherapy, demonstrating that connectivity mapping can identify candidate repurposing opportunities across multiple cancer types, including NSCLC, by matching tumor-specific expression signatures against the perturbation library [PMID: 37068308]. However, a persistent limitation of CMap-based approaches is their reliance on transcriptomic signatures as proxies for therapeutic effect: gene expression changes induced by a drug in vitro do not necessarily predict clinical efficacy, and the relationship between signature reversal and tumor regression is correlative rather than mechanistic.
Tanoli and colleagues provided a comprehensive review of the drug repurposing field, cataloging the major computational strategies — including network-based, structure-based, signature-based, and knowledge graph approaches — and evaluating their track record in generating clinically validated repurposing candidates [PMID: 40102635]. Their assessment is candid: while computational repurposing has generated thousands of hypotheses, the number that have progressed through clinical validation to regulatory approval remains small. The attrition rate reflects not a failure of computational methods per se but the inherent difficulty of predicting in vivo therapeutic efficacy from in silico or in vitro data. The field needs more disciplined validation pipelines — and, crucially, more clinical champions willing to invest trial resources in repurposing candidates rather than novel chemical entities.
Gene-Drug Interaction Databases and Network Pharmacology
The infrastructure supporting computational drug repurposing has expanded considerably with the development of curated gene-drug interaction databases. The Drug Gene Interaction Database (DGIdb), now in its fifth iteration, aggregates data from over 40 source databases to catalog known and predicted interactions between genes and drugs, providing a queryable resource for identifying existing compounds that target genes of interest [PMID: 37953380]. DGIdb 5.0 integrates druggability predictions, enabling researchers to assess not only whether a given lung cancer target has known ligands but also whether structurally related proteins in the same family are druggable — a critical consideration for targets that lack direct pharmacological tools but may be vulnerable to indirect inhibition through pathway-level interventions.
Network pharmacology extends the single-target paradigm by modeling drug effects as perturbations across interconnected molecular networks. MotieGhader and colleagues applied network-based drug repositioning specifically to NSCLC, integrating protein-protein interaction networks with drug-target data to identify compounds predicted to disrupt cancer-associated network modules [PMID: 35676421]. Their approach exemplifies a growing recognition that the therapeutic effect of a drug often derives not from inhibition of a single target but from modulation of a network neighborhood — an insight that aligns with the systems biology perspective but complicates the clean biomarker-target-drug logic that underlies most precision oncology frameworks. Nair and colleagues explored drug combination strategies for NSCLC through computational analysis, identifying synergistic pairings that target complementary vulnerabilities within the tumor signaling network [PMID: 37380628]. Combination repurposing — using two or more existing drugs whose individual effects are modest but whose combined action is synergistic — represents an underexplored strategy that could substantially expand the clinical utility of the existing pharmacopoeia.
Pharmacogenomics and Liquid Biopsy-Guided Repurposing
The integration of pharmacogenomic data with real-time tumor monitoring through liquid biopsy creates a new paradigm for adaptive drug repurposing. Kolesar and colleagues described the intersection of liquid biopsy technology and pharmacogenomics, arguing that circulating tumor DNA (ctDNA) profiling can identify emergent resistance mutations in real time, enabling clinicians to pivot to alternative agents — including repurposed compounds — before clinical progression becomes apparent [PMID: 35209919]. This vision of dynamic, biomarker-guided therapeutic adaptation is conceptually powerful but practically demanding: it requires not only sensitive ctDNA assays but also a curated knowledge base linking specific resistance genotypes to alternative therapeutic options, many of which may involve off-label or repurposed agents. The computational infrastructure for this kind of real-time decision support exists in prototype form but has not been validated in prospective clinical studies.
Synthetic Lethality: The Next Frontier
The most scientifically compelling direction for drug repurposing in NSCLC lies in the exploitation of synthetic lethal interactions — genetic vulnerabilities that emerge only in the context of specific tumor-intrinsic alterations. The concept is borrowed from yeast genetics: two genes are synthetically lethal if loss of either alone is tolerated but loss of both is fatal. In cancer, synthetic lethality offers a strategy for targeting tumors through their own genetic defects — converting an oncogenic alteration from a survival advantage into a therapeutic liability. Leung and colleagues provided an early computational framework for identifying synthetic lethal partners in cancer, demonstrating that large-scale loss-of-function screens can be integrated with genomic data to nominate druggable synthetic lethal targets [PMID: 27686855]. The paradigmatic example is the sensitivity of BRCA-mutant tumors to PARP inhibition, and the extension of this logic to NSCLC-relevant alterations has begun to yield actionable results.
Long and colleagues demonstrated that tumors with loss of LKB1 (STK11), a frequent co-mutation in KRAS-driven NSCLC that confers resistance to immunotherapy, exhibit vulnerability to PARP inhibition through a mechanism linked to impaired DNA damage repair [PMID: 36512628]. This finding is of particular translational significance because LKB1 loss defines a patient population with limited therapeutic options — KRAS/LKB1 co-mutant NSCLC responds poorly to both checkpoint inhibitors and conventional chemotherapy — and PARP inhibitors are already approved drugs with well-characterized safety profiles. The repurposing of an existing drug class to a new indication defined by a specific genomic context exemplifies the kind of synthetic lethality-guided repurposing that this theme should prioritize. Postel-Vinay and colleagues reviewed the broader landscape of DNA repair deficiencies in cancer and their therapeutic exploitation, identifying additional repair pathway vulnerabilities that may be targetable with existing agents [PMID: 22330686].
KRAS: The Repurposing Imperative
KRAS-mutant NSCLC — the single largest molecularly defined subset of lung adenocarcinoma — has historically been considered undruggable, a characterization that persisted until the recent development of KRAS G12C-specific covalent inhibitors. Yet KRAS G12C accounts for only a fraction of KRAS mutations, and patients with G12D, G12V, and other KRAS variants remain without direct targeted options. Wood and colleagues reviewed the therapeutic landscape for KRAS-mutant cancers, emphasizing that indirect strategies — including MEK inhibition, SHP2 inhibition, and combination approaches targeting KRAS effector pathways — remain essential for the majority of KRAS-mutant patients [PMID: 27100819]. Many of the agents targeting these downstream effectors are approved or in advanced clinical development for other indications, making them natural candidates for repurposing to KRAS-mutant NSCLC.
Rosell and colleagues provided a broad overview of the lung cancer therapeutic landscape circa 2014, documenting the state of targeted therapy development and identifying unmet needs that persist today [PMID: 25533943]. Notably, several of the gaps identified in that review — particularly for KRAS-mutant and STK11-mutant tumors — remain open, underscoring the slow pace of translational progress in these genotypes. Linardou and colleagues conducted a meta-analysis examining the relationship between KRAS mutations and resistance to EGFR-targeted therapy, establishing that KRAS mutation status is a negative predictive biomarker for EGFR tyrosine kinase inhibitor response [PMID: 18804418]. This finding, while focused on exclusion rather than inclusion, has repurposing implications: it directs therapeutic attention toward alternative pathways in KRAS-mutant tumors and away from agents whose mechanism of action is antagonized by constitutive RAS activation.
Germline Variants and Population-Level Repurposing Opportunities
The integration of germline genetic data with drug repurposing strategies opens a population-level dimension that is rarely explored in the lung cancer literature. McKay and colleagues reported genome-wide association study (GWAS) results for lung cancer susceptibility, identifying germline variants in regions harboring pharmacologically relevant genes [PMID: 28604730]. While GWAS hits are primarily relevant to risk prediction rather than therapeutic selection, the co-localization of lung cancer risk loci with targets of existing drugs creates hypothesis-generating opportunities for chemoprevention or early intervention using repurposed agents. This application — using drugs developed for other conditions to prevent or delay lung cancer in genetically at-risk populations — is conceptually distinct from the repurposing of drugs for treatment of established tumors and has received almost no attention in the computational lung cancer literature.
The Path Forward: From Drug-Matching Frameworks to Mechanistic Repurposing
The current evidence base for drug repurposing in NSCLC is dominated by what might be termed drug-matching frameworks (DMF): computational approaches that match existing drugs to tumor profiles based on expression signatures, network proximity, or gene-drug interaction databases. These approaches are valuable for hypothesis generation but suffer from high false-positive rates and limited mechanistic resolution. The field must evolve beyond DMF toward mechanistic repurposing strategies anchored in synthetic lethality, metabolic vulnerability, and context-dependent drug sensitivity. The key ingredients for this evolution are already available: large-scale functional genomics screens (CRISPR-Cas9 dependency maps), pharmacogenomic databases linking drug sensitivity to molecular features, and clinical outcome data from molecularly profiled patient cohorts. What is missing is the deliberate integration of these resources into repurposing-specific computational pipelines and, crucially, the clinical trial infrastructure to validate computationally nominated repurposing candidates. With only 24 Tier B papers, this theme is ripe for expansion — and the synthetic lethality paradigm, particularly in the LKB1-loss and DNA repair-deficient contexts, offers the most immediate translational opportunities.
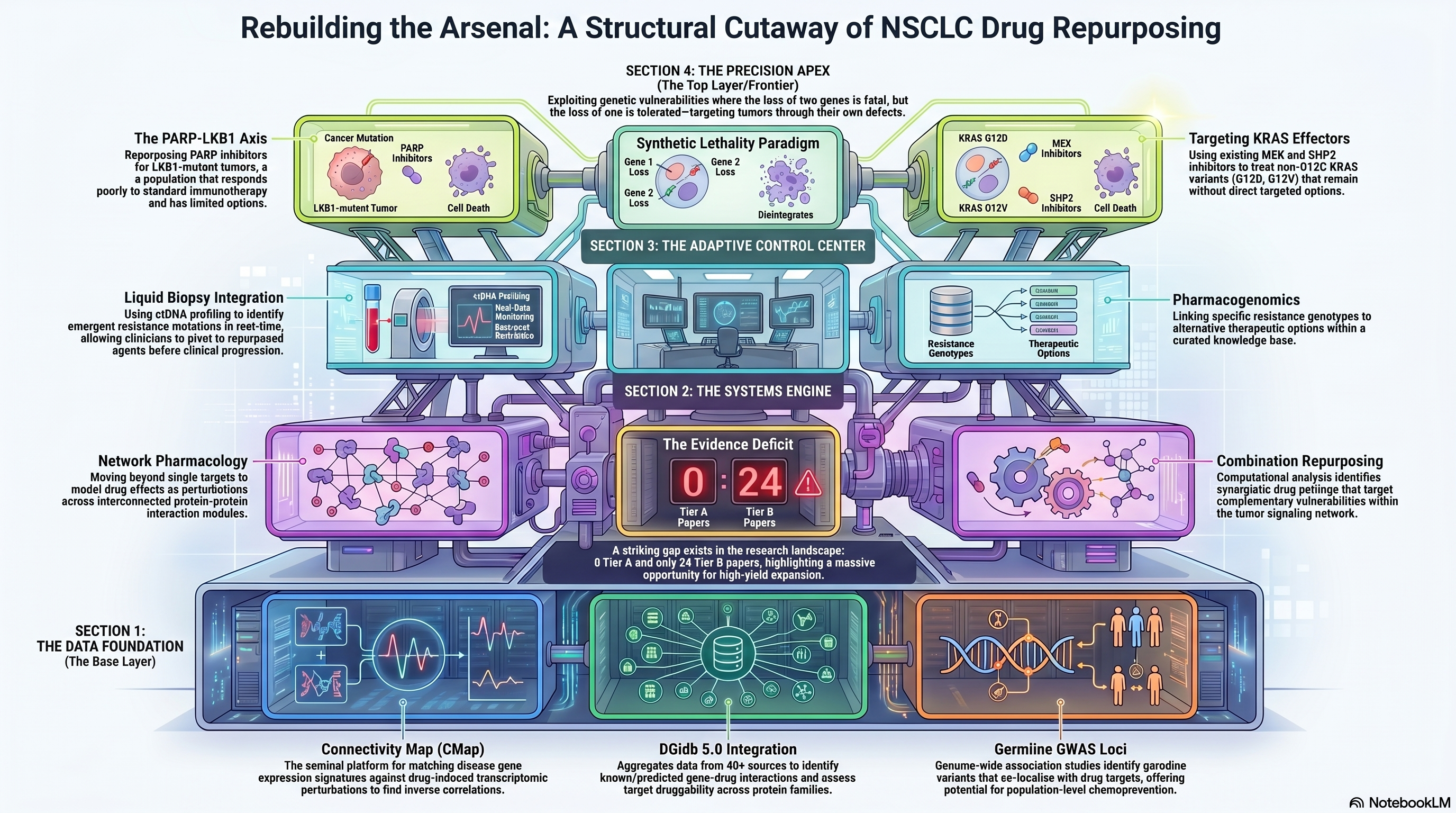 Infographic generated via NotebookLM from the chapter source material.
Infographic generated via NotebookLM from the chapter source material.
Implications for the Manuscript
This theme carries the thinnest evidence base in the entire corpus (0 Tier A, only 24 Tier B), and the manuscript should explicitly flag this deficit as a finding in its own right. The review should argue that drug repurposing in NSCLC has been underserved by the research community relative to its translational potential, and that the field must move beyond traditional drug-matching frameworks toward synthetic lethality-based strategies that exploit tumor-specific genetic vulnerabilities with existing pharmacological agents. Key recommendations include: (1) systematic exploitation of CRISPR dependency data to nominate synthetic lethal partners for common NSCLC alterations (KRAS, STK11, KEAP1); (2) integration of liquid biopsy-guided pharmacogenomics with repurposing knowledge bases for adaptive therapy; (3) dedicated clinical trial designs (e.g., repurposing-focused basket trials) to accelerate validation of computationally nominated candidates; and (4) expansion of the evidence base from the current 24 Tier B papers to a level commensurate with the theme's clinical importance. The PARP-LKB1 axis represents the most mature synthetic lethality opportunity and should be highlighted as a model for future repurposing efforts.