Lung Cancer in Never-Smokers
"Never-smoker lung cancer is not the absence of a cause; it is a distinct disease sculpted by its own mutational forces, molecular subtypes, and epidemiological architecture."
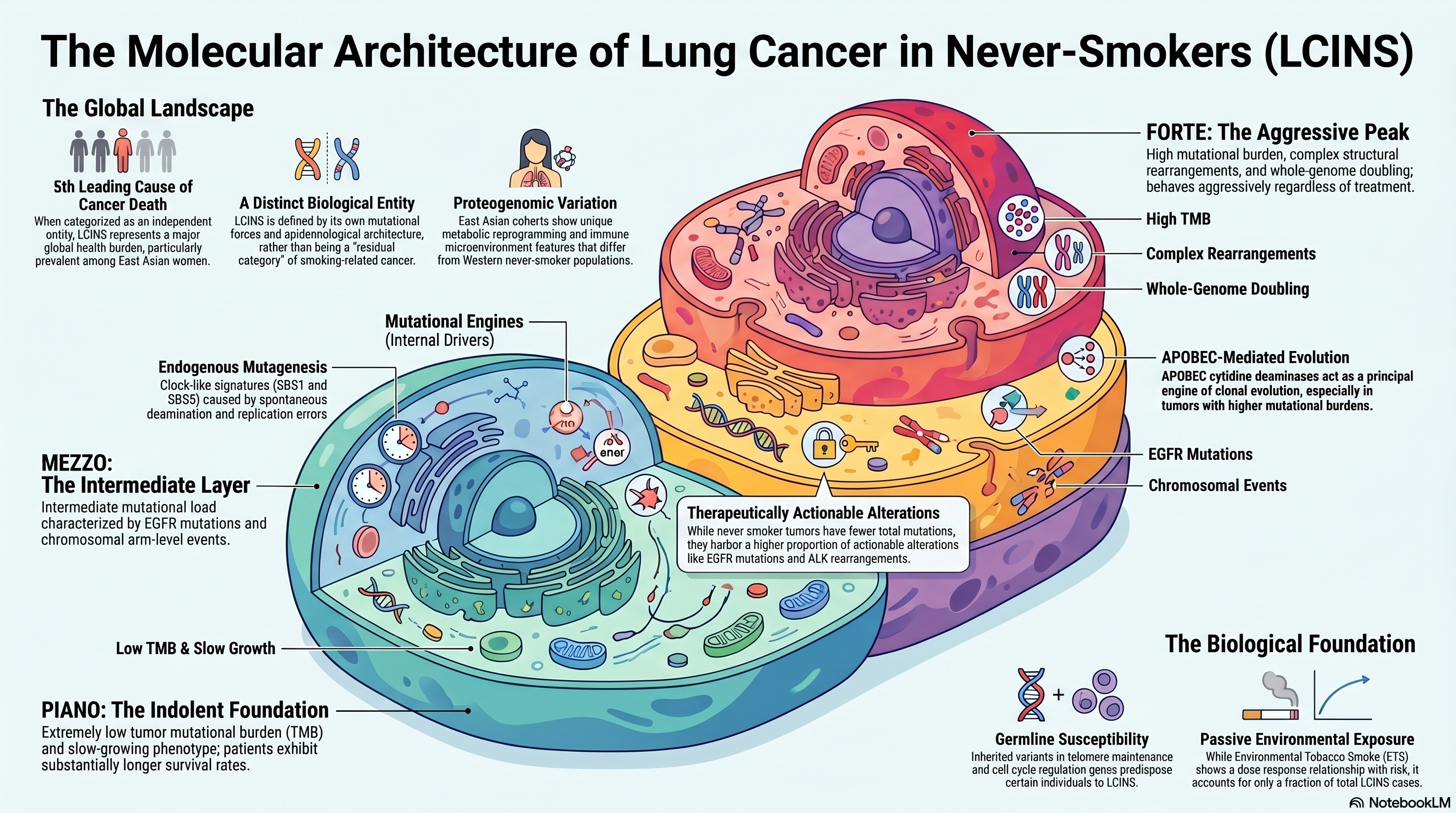 Infographic generated via NotebookLM from the chapter source material.
Infographic generated via NotebookLM from the chapter source material.
Landscape stats: 2,689 papers mapped to this theme (2 Tier A, 76 Tier B). The literature spans molecular subtyping, whole-genome characterization, environmental risk factors, and therapeutic sensitivity, collectively establishing lung cancer in never-smokers (LCINS) as a biologically and clinically distinct entity.
The Scale of a Neglected Problem
Lung cancer in individuals who have never smoked tobacco constitutes the fifth leading cause of cancer death worldwide when considered as an independent entity. Despite this burden, LCINS was historically treated as an epidemiological curiosity, a residual category defined by the absence of the dominant risk factor. Sun and colleagues provided one of the first comprehensive reviews of the never-smoker lung cancer literature, synthesizing evidence on incidence patterns, molecular features, and potential etiological agents, and made the case that LCINS deserved recognition as a disease category in its own right rather than a statistical footnote [PMID: 17882278]. Subramanian and colleagues reinforced this position by reviewing the emerging molecular genetics of never-smoker tumors, documenting the high prevalence of EGFR mutations and ALK rearrangements relative to smoker-associated tumors and arguing that these molecular distinctions had direct therapeutic implications [PMID: 18598932]. The conceptual reframing of LCINS as a distinct entity rather than a variant of smoking-associated lung cancer was a necessary precondition for the genomic investigations that followed.
The Genomic Taxonomy: Piano, Mezzo, and Forte
The landmark study by Zhang and colleagues applied whole-genome sequencing to a large cohort of never-smoker lung adenocarcinomas and identified three molecular subtypes, evocatively named piano, mezzo, and forte, based on the intensity of their mutational burden [PMID: 34493867]. The piano subtype was characterized by an extremely low tumor mutational burden, frequent EGFR driver mutations, a slow-growing indolent phenotype, and activation of progenitor cell programs. The mezzo subtype exhibited intermediate mutational load with enrichment for EGFR mutations alongside concurrent chromosomal arm-level events. The forte subtype, the most aggressive, displayed high mutational burden, complex structural rearrangements, and whole-genome doubling, resembling smoker-associated tumors in its genomic complexity despite arising in the absence of tobacco exposure. This tripartite classification provided the first coherent framework for understanding the biological heterogeneity within LCINS and explained, at least in part, the variable clinical behavior that had long puzzled clinicians. The piano-mezzo-forte taxonomy also carried prognostic implications: patients with piano subtype tumors exhibited substantially longer survival, consistent with their indolent biology, while forte subtype tumors behaved aggressively regardless of treatment.
Mutational Processes and Endogenous Mutagenesis
If tobacco smoke is not the mutational driver in LCINS, what forces shape the somatic genome of these tumors? Diaz-Gay and colleagues addressed this question through a systematic analysis of mutational signatures across never-smoker and ever-smoker lung cancers, identifying distinct mutagenic forces operative in each group [PMID: 40604281]. Never-smoker tumors were dominated by clock-like signatures attributed to spontaneous deamination of methylated cytosines (SBS1) and replication-associated errors (SBS5), with additional contributions from APOBEC-mediated mutagenesis (SBS2 and SBS13). The APOBEC signature was particularly notable because it was enriched in tumors with higher mutational burden, suggesting that endogenous mutagenesis by APOBEC cytidine deaminases might serve as the principal engine of clonal evolution in a subset of LCINS. Govindan and colleagues had earlier compared the mutational landscapes of smoker and never-smoker tumors using exome sequencing, reporting that never-smoker tumors harbored fewer total mutations but a higher proportion of therapeutically actionable alterations, a finding with direct clinical relevance [PMID: 22980976]. The convergent finding across these studies was that LCINS genomics is defined not by random mutagenesis but by specific, identifiable mutational processes that impose structure on the somatic genome.
Proteogenomic Characterization in East Asian Cohorts
The epidemiology of LCINS is strikingly non-uniform across populations, with the highest incidence observed in East Asian women, a pattern that has driven several large-scale molecular profiling efforts in Asian cohorts. Chen and colleagues conducted a proteogenomic analysis of lung adenocarcinomas from East Asian never-smokers, integrating whole-genome sequencing, RNA sequencing, and quantitative proteomics to construct a multi-omic portrait of these tumors [PMID: 32649875]. Their analysis identified proteomic subtypes that only partially overlapped with transcriptomic classifications, revealing post-transcriptional regulatory layers that shaped functional tumor phenotypes. Notably, tumors from East Asian never-smokers exhibited distinctive metabolic reprogramming and immune microenvironment features compared with Western cohorts, raising the question of whether population-specific genetic backgrounds or environmental exposures contributed to these differences. The proteogenomic data also nominated several candidate therapeutic vulnerabilities not apparent from genomic data alone, including dependencies on specific metabolic pathways and protein quality control mechanisms.
Clonal Evolution and Tumor Heterogeneity
Understanding how LCINS tumors evolve over time is essential for designing durable therapeutic strategies. Frankell and colleagues, through the TRACERx consortium, characterized the evolutionary trajectories of lung cancers with multi-region and longitudinal sampling, providing insights into the clonal dynamics of tumors including those arising in never-smokers [PMID: 37046096]. Their work demonstrated that never-smoker tumors, despite their lower overall mutational burden, could exhibit significant intratumoral heterogeneity driven by chromosomal instability and parallel evolution of subclonal populations. Chabon and colleagues extended the evolutionary perspective by examining the genomic features of circulating tumor DNA (ctDNA) in patients with early-stage NSCLC, including never-smokers, and found that specific genomic features detectable in ctDNA could predict disease recurrence [PMID: 32269342]. The ctDNA findings were particularly relevant for never-smokers with piano-subtype tumors, where the challenge of identifying the minority of patients destined for recurrence among a largely indolent population is a pressing clinical problem.
Functional Genomics and Pathway Dependencies
Beyond cataloguing mutations, functional genomic approaches have begun to reveal the pathway dependencies that sustain never-smoker tumors. Hamouz and colleagues employed CRISPR-based functional genomic screens to identify genes essential for the survival of EGFR-mutant lung cancer cells, the predominant genotype in LCINS [PMID: 37686122]. Their screens identified both expected dependencies, confirming the central role of EGFR signaling, and unexpected vulnerabilities in RNA processing and chromatin remodeling pathways that could serve as targets for combination therapy. Yano and colleagues characterized the clinical and molecular features of non-smoking NSCLC in a Japanese cohort, documenting the high frequency of actionable mutations and the favorable response rates to targeted therapies [PMID: 21562939]. Janne and colleagues reported on the activity of dacomitinib, a pan-HER inhibitor, in advanced NSCLC, with subgroup analyses demonstrating particular efficacy in the EGFR-mutant, never-smoker population [PMID: 25456362]. Taken together, the functional genomics and clinical trial data support a model in which never-smoker tumors are characterized by deep dependence on a limited number of oncogenic drivers, creating therapeutic opportunities but also vulnerability to resistance through bypass pathway activation.
Germline Susceptibility and Inherited Risk
The clustering of LCINS in specific populations and families has motivated investigations into germline genetic susceptibility. Li and colleagues conducted a genome-wide association study (GWAS) focused on never-smoker lung cancer cases and identified susceptibility loci at chromosomal regions not previously associated with smoking-related lung cancer [PMID: 20304703]. The identified variants mapped to genes involved in telomere maintenance and cell-cycle regulation, suggesting that inherited variation in genomic stability pathways may predispose to tumorigenesis in the absence of exogenous mutagens. The GWAS findings complemented family-based studies documenting elevated lung cancer risk among first-degree relatives of never-smoker patients, collectively arguing that LCINS has a genetic architecture distinct from tobacco-associated lung cancer.
Environmental Tobacco Smoke and Passive Exposure
Although LCINS is defined by the absence of active tobacco use, passive exposure to environmental tobacco smoke (ETS) has been consistently associated with elevated risk. Fontham and colleagues conducted a multicenter case-control study of ETS exposure and lung cancer risk in never-smoking women, reporting a dose-response relationship between cumulative household and workplace ETS exposure and lung cancer incidence [PMID: 8196118]. The Fontham study was influential in establishing the causal link between passive smoking and lung cancer in never-smokers, contributing to subsequent public health interventions including indoor smoking bans. However, ETS exposure accounts for only a fraction of LCINS cases, and the majority of never-smoker lung cancer patients report minimal or no passive smoke exposure, reinforcing the need to identify additional etiological agents.
Implications for the Manuscript
The never-smoker lung cancer literature provides several essential threads for the manuscript. First, the Zhang piano-mezzo-forte taxonomy should be presented as the organizing framework for LCINS biology, because it reconciles the apparent heterogeneity of clinical behavior with underlying molecular subtypes and has direct prognostic relevance. Second, the mutational signature analyses from Diaz-Gay and Govindan establish that endogenous mutagenic processes, particularly APOBEC activity, are the dominant shapers of the LCINS genome, a finding that should be discussed in the context of therapeutic implications such as sensitivity to DNA damage-directed agents. Third, the proteogenomic data from Chen and colleagues illustrate the importance of protein-level analysis in populations where genomic data alone may miss functional complexity, and the manuscript should highlight the population-specific nature of LCINS biology. Fourth, the GWAS findings from Li and colleagues support the argument that germline risk profiling could eventually be incorporated into screening strategies for never-smokers, a population currently excluded from most low-dose CT screening guidelines despite bearing a significant lung cancer burden. The manuscript should argue that LCINS represents a paradigm case in which molecular subtyping, rather than smoking history, should drive clinical decision-making.