Theme 9 — Immune Biomarkers and Immunotherapy
"The immune system does not see cancer as foreign — it sees cancer as self, badly edited."
Corpus snapshot: 6,459 papers · 9 Tier A · 679 Tier B
The Immunogenomic Turn in Lung Cancer
The advent of immune checkpoint inhibitors transformed the treatment landscape for non-small cell lung cancer (NSCLC), but their clinical deployment exposed a profound knowledge gap: only a minority of patients derive durable benefit, and the biological determinants of response remain incompletely understood. The field's initial attempt to bridge this gap centered on a single analyte — tumor mutational burden (TMB) — catalyzed by the landmark observation that high nonsynonymous mutation load correlated with objective response and durable clinical benefit in NSCLC patients treated with pembrolizumab [PMID: 25765070]. That study, now among the most cited in immuno-oncology with over 6,500 citations, established a conceptual framework in which somatic mutations generate neoantigens that render tumors visible to adaptive immunity. Yet the intervening decade has revealed that TMB is a necessary but insufficient predictor, and the search for complementary, orthogonal, and integrative biomarkers has become one of the defining research programs of computational lung cancer biology.
From Mutation Counts to Immune Ecosystems
The reductionism inherent in single-analyte biomarkers was forcefully challenged by pan-cancer immunogenomic analyses that classified tumors not by mutation count alone but by the composite architecture of their immune microenvironments. The Cancer Genome Atlas consortium's comprehensive immune landscape analysis stratified over 10,000 tumors into six immune subtypes defined by differences in macrophage and lymphocyte signatures, Th1-to-Th2 cell ratios, extent of intratumoral heterogeneity, and aneuploidy [PMID: 29628290]. Lung adenocarcinoma and squamous cell carcinoma partitioned differentially across these subtypes, with prognostic implications that could not be captured by TMB alone. This pan-cancer taxonomy established that the tumor microenvironment (TME) is not a monolithic entity but a mosaic of distinct ecological states, each with different therapeutic vulnerabilities. Extending this logic, Bagaev and colleagues developed a four-archetype classification of TME subtypes — immune-enriched, immune-enriched fibrotic, immune-desert, and fibrotic — using transcriptomic deconvolution of 29 functional gene expression signatures across thousands of solid tumors [PMID: 34019806]. Their framework demonstrated that TME archetype predicts immunotherapy response across cancer types more reliably than TMB in isolation, and that the stromal and myeloid compartments carry prognostic information at least as important as that derived from T cell infiltration metrics. In NSCLC specifically, the immune-desert and fibrotic subtypes were associated with primary resistance to checkpoint blockade, suggesting that biomarker strategies must account for the non-immune cellular milieu.
Computational Immunophenotyping and the Immunophenoscore
The challenge of operationalizing TME complexity for clinical decision-making prompted the development of computational tools designed to translate bulk transcriptomic data into immunologically interpretable scores. The Immunophenoscore, a machine-learning-derived metric integrating effector cells, immunosuppressive cells, MHC-related molecules, and immunomodulators, demonstrated predictive value for checkpoint inhibitor response across multiple validation cohorts [PMID: 28052254]. Its strength lay in its ability to aggregate signals from antagonistic immune populations — for instance, simultaneously capturing the presence of cytotoxic T lymphocytes and the countervailing influence of regulatory T cells or myeloid-derived suppressor cells. This approach anticipated a broader trend in the field: the recognition that no single cell type determines immunotherapy outcome, and that what matters is the balance of power between pro-inflammatory and immunosuppressive forces within the tumor ecosystem.
The relationship between TMB and immune infiltration itself proved more complex than initially assumed. Ricciuti and colleagues provided critical clarification by demonstrating that TMB and T cell infiltration are partially independent predictors of immunotherapy benefit in NSCLC, and that patients with both high TMB and high CD8+ T cell density derive substantially greater benefit than those with either feature alone [PMID: 35708671]. This interaction effect underscores the intuition that mutations must not only generate neoantigens but that those neoantigens must be presented, recognized, and acted upon by a competent immune effector apparatus. The two-axis model — mutational supply and immune demand — has since become a conceptual anchor for composite biomarker development.
Antigen Presentation, Immunopeptidomes, and the HLA Interface
If mutations supply neoantigens and T cells provide effector function, then the major histocompatibility complex (MHC) serves as the critical intermediary. Recent work has brought the immunopeptidome — the repertoire of peptides actually presented on MHC molecules — into sharper focus as a determinant of immunotherapy response. Kraemer and colleagues mapped the immunopeptidome across NSCLC tumors and demonstrated that the diversity and composition of presented peptides varies substantially between patients, driven in part by HLA genotype and tumor-specific alterations in antigen processing machinery [PMID: 37127787]. This work elevated antigen presentation from a theoretical bottleneck to an empirically measurable variable. In a complementary line of investigation, Lester and colleagues uncovered a role for fucosylation of HLA-DRB1 in modulating peptide presentation and immune recognition, revealing a post-translational layer of regulation that standard genomic analyses entirely miss [PMID: 36690875]. These findings collectively argue that biomarker strategies focused exclusively on mutational profiles or bulk transcriptomic signatures will fail to capture the mechanistic complexity governing whether a neoantigen is seen, ignored, or actively tolerated by the immune system.
Beyond Conventional T Cells: Gamma-Delta Cells, Retention Signals, and Microbiome Modifiers
The dominant narrative of anti-tumor immunity in NSCLC has centered on conventional alpha-beta T cells, particularly CD8+ cytotoxic lymphocytes. However, emerging evidence points to the functional importance of unconventional lymphocyte populations that operate through distinct recognition mechanisms. De Vries and colleagues characterized gamma-delta T cells within the NSCLC tumor microenvironment, demonstrating that these MHC-unrestricted lymphocytes exhibit both pro-tumorigenic and anti-tumorigenic phenotypes depending on their subtype and activation context [PMID: 36631610]. The duality of gamma-delta T cell function — capable of direct cytotoxicity via NKG2D but also of immunosuppressive cytokine production — complicates therapeutic strategies that aim to harness innate-like immunity but highlights an underexplored axis for biomarker development. Wang and colleagues investigated the mechanisms governing T cell retention within NSCLC tumors, identifying molecular programs that determine whether infiltrating lymphocytes persist in the tumor or are expelled to the periphery [PMID: 40341231]. This retention biology intersects with the concept of tissue-resident memory T cells, a population increasingly recognized as central to durable anti-tumor immunity.
The microbiome has emerged as an unexpected modifier of immunotherapy response, though the mechanistic pathways remain actively debated. Stein-Thoeringer and colleagues demonstrated that gut microbiome composition influences outcomes in patients receiving immune checkpoint inhibitors, with specific bacterial taxa associated with enhanced or diminished therapeutic benefit [PMID: 36914893]. Intriguingly, Ferrari and colleagues extended the microbiome-immunity axis by showing that microbial metabolites can influence HLA-mediated antigen presentation, establishing a potential mechanistic link between commensal organisms and adaptive anti-tumor immunity [PMID: 37738976]. These findings suggest that the predictive biomarker space extends beyond the tumor and its immediate microenvironment to encompass host-level ecological variables that are rarely captured in standard clinical workflows.
Immune Evasion and Therapeutic Resistance: p53 and Beyond
The flip side of biomarker discovery is understanding the mechanisms by which tumors evade immune destruction despite nominally favorable biomarker profiles. TP53 mutations, present in roughly half of NSCLC cases, have long been associated with genomic instability and elevated TMB, yet their relationship to immunotherapy outcome is paradoxically inconsistent. Zhu and colleagues elucidated a mechanism by which specific p53 mutations actively suppress anti-tumor immunity through modulation of innate immune sensing pathways, providing a molecular explanation for immunotherapy resistance in tumors that would otherwise be predicted to respond based on their mutational load [PMID: 37774698]. This work illustrates a critical principle: the same oncogenic mutation can simultaneously increase neoantigen production and cripple the immune response required to exploit those neoantigens. The net effect on immunotherapy sensitivity depends on which of these opposing forces dominates, and predicting this balance remains a formidable computational challenge.
Emerging Therapeutic Modalities: Vaccines, Bispecifics, and Engineered Immunity
The biomarker landscape is not solely shaped by diagnostic need; therapeutic innovation creates new demands for predictive tools. Dendritic cell (DC) vaccines represent a strategy to prime adaptive immunity against tumor-specific antigens, and Li and colleagues reported advances in DC vaccine design that leverage neoantigen prediction pipelines to personalize vaccine composition for individual NSCLC patients [PMID: 39613742]. The success of such approaches depends critically on accurate neoantigen identification and HLA binding prediction, tying vaccine development directly to the immunopeptidomic analyses discussed above. On the antibody engineering front, bispecific antibodies targeting PD-1 and a second immune checkpoint or co-stimulatory molecule represent the next generation of checkpoint-based therapy. Zeng and colleagues described a PD-1-targeting bispecific construct designed to simultaneously engage two immunomodulatory pathways, with preclinical evidence of enhanced anti-tumor efficacy compared to monospecific PD-1 blockade [PMID: 41398162]. The biomarker implications are significant: if combination immunomodulation becomes standard, composite biomarkers will need to capture the status of multiple immune axes simultaneously, further increasing the dimensionality of the prediction problem.
The Integration Challenge: From Individual Markers to Systems-Level Prediction
The trajectory of immune biomarker research in NSCLC is one of increasing dimensionality. What began as a single-variable question — does TMB predict response? — has expanded into a multidimensional optimization problem encompassing mutation burden, neoantigen quality, MHC diversity, immunopeptidome composition, TME architecture, T cell phenotype and retention, microbiome composition, and tumor-intrinsic immune evasion mechanisms. Each of these axes contributes independently to the prediction of immunotherapy outcome, and their interactions — some synergistic, some antagonistic — create a combinatorial space that exceeds the capacity of conventional biomarker frameworks. The field is now at an inflection point where machine learning and multi-omic integration are not luxuries but necessities, and where the design of clinical trials must accommodate biomarker complexity rather than reduce it to binary cutoffs. The challenge is not merely technical but epistemological: the immune system is a dynamic, adaptive network, and static biomarker measurements at a single time point capture only a frozen snapshot of a continuously evolving interaction between host and tumor.
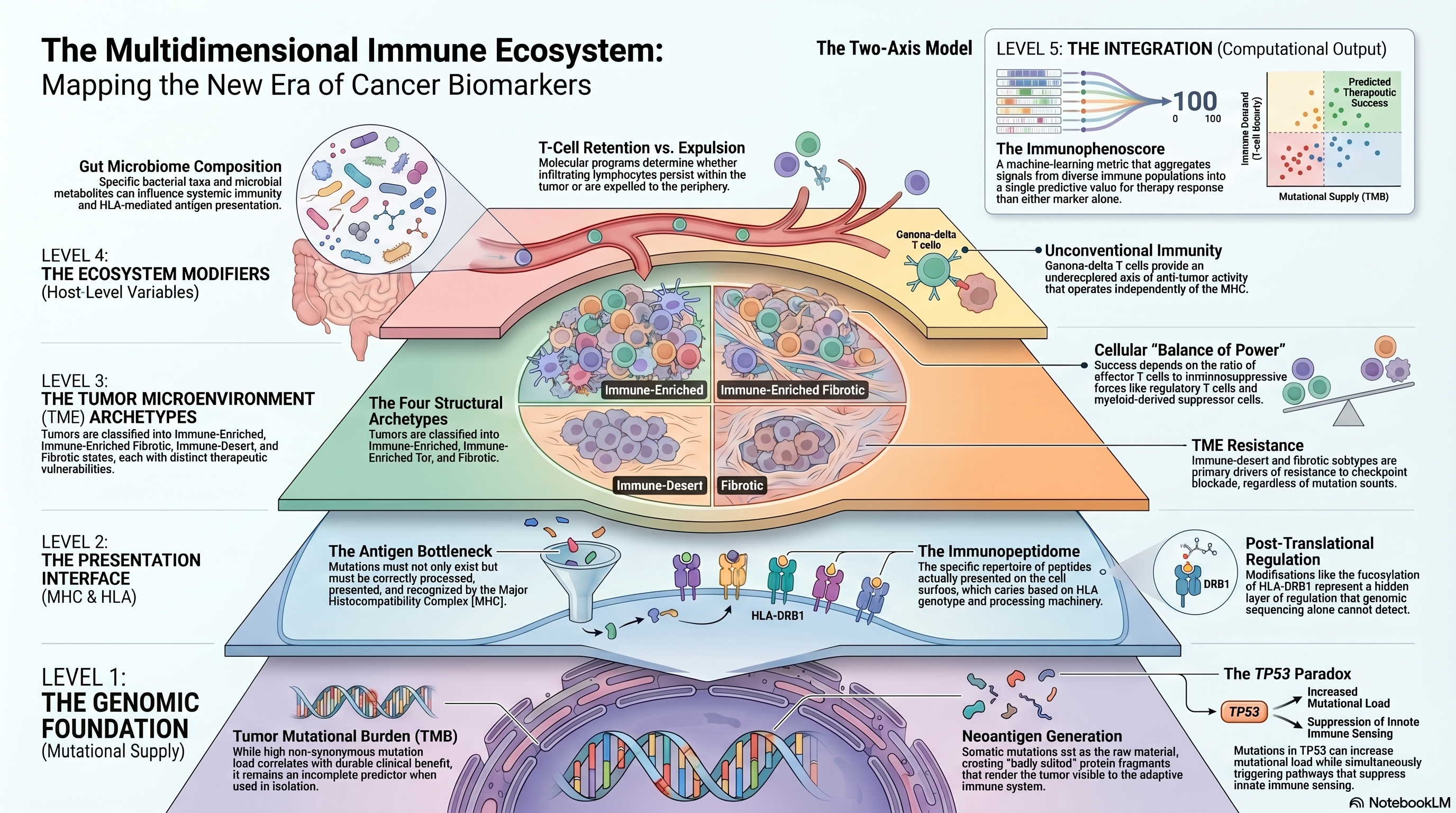 Infographic generated via NotebookLM from the chapter source material.
Infographic generated via NotebookLM from the chapter source material.
Implications for the Manuscript
This theme demonstrates that the immune biomarker field has matured far beyond TMB as a standalone predictor. The manuscript should emphasize three key transitions: (1) from single-analyte to composite, ecosystem-level biomarkers that integrate TME architecture with mutational profiles; (2) from genomic-only to multi-omic frameworks that incorporate the immunopeptidome, post-translational modifications, and microbiome data; and (3) from static to dynamic biomarker concepts that account for immune evasion mechanisms and T cell retention biology. The 9 Tier A papers in this corpus anchor a narrative of progressive sophistication, while the 679 Tier B papers provide the granular mechanistic detail necessary for a comprehensive review. The manuscript should position composite biomarker development as a high-priority area where computational biology can deliver clinical impact in the near term, while acknowledging that validation in prospective cohorts remains the critical bottleneck.