Theme 10 — The Translational Bottleneck: From Discovery to Delivery
"We do not lack for targets. We lack for equity, infrastructure, and the institutional will to close the gap between what we know and whom we treat."
Corpus snapshot: 6,431 papers · 0 Tier A · 67 Tier B
The Paradox of Abundance
Contemporary lung cancer biology produces genomic, transcriptomic, and proteomic data at a pace that far outstrips the clinical infrastructure required to act on it. The field now commands detailed molecular atlases of tumor heterogeneity, validated therapeutic targets across dozens of driver alterations, and increasingly sophisticated predictive biomarkers — yet the majority of lung cancer patients worldwide never receive molecular testing, and among those who do, actionable findings frequently fail to translate into treatment changes. This theme confronts the translational bottleneck: the systemic, structural, and sociological barriers that prevent precision oncology from reaching the patients who need it most. Notably, this is the only major theme in the corpus with zero Tier A papers, a bibliometric signal that the field's most-cited work has overwhelmingly favored discovery over implementation. The 67 Tier B papers that anchor this theme address disparities, real-world evidence, molecular tumor boards, and the practical mechanics of moving biomarker-matched therapies from academic centers to community practice.
Racial and Socioeconomic Disparities: A Decades-Old Problem
The inequitable distribution of lung cancer outcomes across racial and socioeconomic groups was documented well before the genomic era. Bach and colleagues published a foundational analysis demonstrating significant racial differences in lung cancer treatment and survival, with Black patients receiving surgical resection at lower rates than white patients even after adjustment for stage, comorbidity, and insurance status [PMID: 10519898]. That study, now cited over 540 times, established the empirical basis for a disparity that has proven remarkably resistant to correction over the subsequent quarter century. The persistence of this gap into the precision medicine era is documented by Dutta and colleagues, who showed that disparities in access to genomic testing, clinical trials, and targeted therapies disproportionately affect patients from minority and low-income populations [PMID: 37248397]. The implications for computational oncology are direct: algorithms trained predominantly on data from well-resourced academic centers risk encoding and amplifying existing biases, producing biomarker models that perform well in the populations they were developed in and poorly in the populations that bear the greatest burden of disease.
Li and colleagues conducted an umbrella review synthesizing evidence across multiple systematic reviews on the relationship between socioeconomic status (SES) and lung cancer outcomes, confirming a consistent gradient in which lower SES is associated with later-stage diagnosis, reduced access to guideline-concordant therapy, and worse survival across virtually every healthcare system studied [PMID: 39557933]. The mechanisms driving this gradient are multifactorial — spanning insurance coverage, geographic access to specialized centers, health literacy, occupational exposures, and comorbidity burden — but the net effect is a systematic exclusion of disadvantaged populations from the benefits of precision medicine. Any manuscript claiming to map the landscape of computational lung cancer research must reckon with this exclusion, not as a peripheral concern but as a central failure of the translational pipeline.
Ancestry-Aware Biomarkers and the Diversity Deficit
The genomic architecture of lung cancer varies across ancestral populations in ways that directly affect the performance of biomarker panels and therapeutic selection algorithms. Arora and colleagues highlighted the critical need for ancestry-aware biomarker development, demonstrating that allele frequencies of pharmacogenomically relevant variants, tumor mutational spectra, and driver gene prevalence differ substantially across populations of European, African, East Asian, and South Asian descent [PMID: 39786754]. The practical consequence is that biomarker thresholds calibrated in one population may misclassify patients in another — a problem that is invisible in ethnically homogeneous discovery cohorts but becomes clinically significant when algorithms are deployed globally. The same point applies to germline pharmacogenomic variants that influence drug metabolism, toxicity profiles, and optimal dosing: biomarker panels that ignore ancestry risk both undertreatment and overtreatment in underrepresented populations.
Hofman and colleagues addressed the biomarker implementation gap from a different angle, proposing practical solutions for integrating comprehensive biomarker testing into routine clinical workflows [PMID: 39904223]. Their analysis identified specific barriers — insufficient tissue procurement, fragmented testing pathways, reimbursement obstacles, and lack of standardized reporting formats — that collectively explain why even well-validated biomarkers fail to reach the point of care. The gap between biomarker validation and biomarker utilization is not a knowledge problem; it is an engineering and policy problem, and computational solutions that do not address implementation logistics will remain confined to academic publications.
Comprehensive Genomic Profiling in the Real World
The promise of comprehensive genomic profiling (CGP) — sequencing hundreds of cancer-related genes in a single clinical assay — rests on the assumption that broad molecular characterization will increase the proportion of patients matched to effective therapies. Saito and colleagues evaluated CGP in a real-world clinical setting, reporting on the rates of actionable findings, treatment changes, and clinical outcomes among patients who received CGP as part of routine care rather than within a clinical trial [PMID: 41495408]. Their findings revealed both the potential and the limitations of real-world CGP: while a substantial fraction of patients harbored potentially actionable alterations, the rate at which these findings translated into treatment modifications was lower than anticipated, constrained by drug availability, patient fitness, and clinical judgment. This real-world evidence serves as a corrective to the optimistic framing of precision oncology trials, in which patient selection and protocol adherence inflate the apparent clinical utility of genomic testing.
The SAFIR02-Lung trial, reported by Besse and colleagues, provided a more structured assessment of biomarker-driven therapy in advanced NSCLC. This randomized trial tested whether maintenance therapy matched to genomic alterations identified by high-throughput molecular profiling improved progression-free survival compared to standard maintenance chemotherapy [PMID: 38351187]. The results were sobering: while individual patients derived benefit from matched therapy, the overall trial did not demonstrate a statistically significant advantage for the genomics-guided arm, raising difficult questions about the design and interpretation of umbrella and basket trials in lung cancer. The trial's contribution lies less in its topline result than in its granular documentation of the logistical challenges — screening failures, tissue insufficiency, turnaround time, molecular heterogeneity — that attenuate the real-world impact of precision oncology.
Molecular Tumor Boards: The Human-in-the-Loop
Molecular tumor boards (MTBs) represent the institutional mechanism by which genomic findings are translated into clinical recommendations. These multidisciplinary forums — typically comprising medical oncologists, pathologists, molecular biologists, bioinformaticians, and clinical geneticists — review complex genomic reports and synthesize recommendations that account for clinical context, evidence quality, and drug access. Tsimberidou and colleagues provided a comprehensive review of MTB structure, function, and outcomes, documenting the growing evidence that MTB-guided therapy improves response rates and survival compared to physician's choice in patients with advanced cancers [PMID: 37845306]. However, the scalability of MTBs remains a fundamental concern: each case requires substantial expert time, and the supply of molecular oncology expertise is insufficient to support MTBs at community cancer centers where the majority of lung cancer patients receive care.
Kato and colleagues described the N-of-One MTB model, an approach that combines automated variant annotation with expert clinical interpretation to deliver personalized treatment recommendations at scale [PMID: 33009371]. This hybrid model — part computational, part human — represents a pragmatic compromise between the interpretive rigor of traditional MTBs and the throughput required for population-level implementation. Tamborero and colleagues developed the MTB Portal, a computational platform designed to standardize variant interpretation and facilitate knowledge sharing across institutions [PMID: 35221333]. The portal enables participating centers to access curated evidence for variant-drug associations, reducing redundant expert effort and harmonizing recommendations. Together, these initiatives suggest that the MTB of the future will be a cyber-physical system in which machine interpretation handles routine variants while human expertise is reserved for genuinely ambiguous cases.
Treatment Complexity and Decision Support
The expanding menu of targeted therapies and immunotherapies for NSCLC has created a decision-making problem that exceeds the cognitive bandwidth of individual clinicians. Meyer and colleagues reviewed the current treatment landscape for NSCLC, cataloging the approved therapies, biomarker requirements, sequencing considerations, and resistance mechanisms that clinicians must navigate for each patient [PMID: 39121882]. The complexity is staggering: a single patient with EGFR-mutant NSCLC now faces a decision tree that includes first-line osimertinib, potential amivantamab-lazertinib combinations, consideration of resistance mechanisms (C797S, MET amplification, histologic transformation), and salvage options that depend on the specific resistance genotype detected at progression. Clinical decision support tools — ranging from simple rule-based algorithms to machine learning systems trained on real-world outcome data — are not optional enhancements but essential infrastructure for managing this complexity at scale.
Reardon and colleagues described the convergence of machine learning and genomics as a framework for clinical decision support, arguing that the integration of multi-omic data with clinical variables through ML architectures can identify treatment-response patterns that are invisible to human cognition [PMID: 41478861]. While the promise is substantial, the validation requirements are formidable: clinical ML models must demonstrate not only discriminative accuracy but also calibration, fairness across demographic groups, and interpretability sufficient for clinician trust.
Global Access and the Trials That Should Exist
The translational bottleneck is sharpest in low- and middle-income countries, where neither the diagnostic infrastructure for molecular profiling nor the therapeutic agents identified by profiling are reliably available. Xu and colleagues reported on icotinib, a domestically developed EGFR tyrosine kinase inhibitor in China, which was designed and tested specifically to address the access gap for targeted therapy in a population with high EGFR mutation prevalence [PMID: 35862035]. The development of icotinib illustrates both the potential and the limitations of regional pharmaceutical innovation as a strategy for closing the access gap: while it expanded EGFR-targeted therapy to millions of patients who might not otherwise have received it, the underlying model of biomarker-driven treatment selection still requires molecular testing infrastructure that remains unevenly distributed.
Arrieta and colleagues conducted a trial of pembrolizumab in Latin American NSCLC patients, providing critical data on immunotherapy outcomes in a population underrepresented in registration trials [PMID: 32271354]. Their findings highlighted both similarities and differences in response patterns compared to trials conducted predominantly in North American, European, and East Asian populations, reinforcing the argument that global biomarker validation requires global clinical trial participation. The computational corollary is clear: predictive models trained on data from a narrow demographic base cannot be assumed to generalize, and the field must invest in diverse training datasets as deliberately as it invests in algorithmic innovation.
The Institutional Ecology of Translation
The translational bottleneck is not a single barrier but an ecosystem of interlocking constraints: insufficient tissue for molecular testing, fragmented electronic health records that impede real-world evidence generation, regulatory frameworks that lag behind technological capability, reimbursement policies that disincentivize comprehensive testing, workforce shortages in molecular pathology and bioinformatics, and clinical trial designs that exclude the patients most likely to benefit from precision approaches. Each of these constraints has been individually documented, but their compound effect — the multiplicative attrition that reduces the fraction of patients who benefit from precision oncology — has received insufficient attention. The zero Tier A papers in this theme is itself a finding: the field's citation economy rewards discovery over implementation, novelty over equity, and molecular elegance over clinical logistics. A manuscript that aspires to guide future research investment must explicitly name this imbalance and advocate for the computational infrastructure, clinical trial reforms, and policy interventions needed to close the gap.
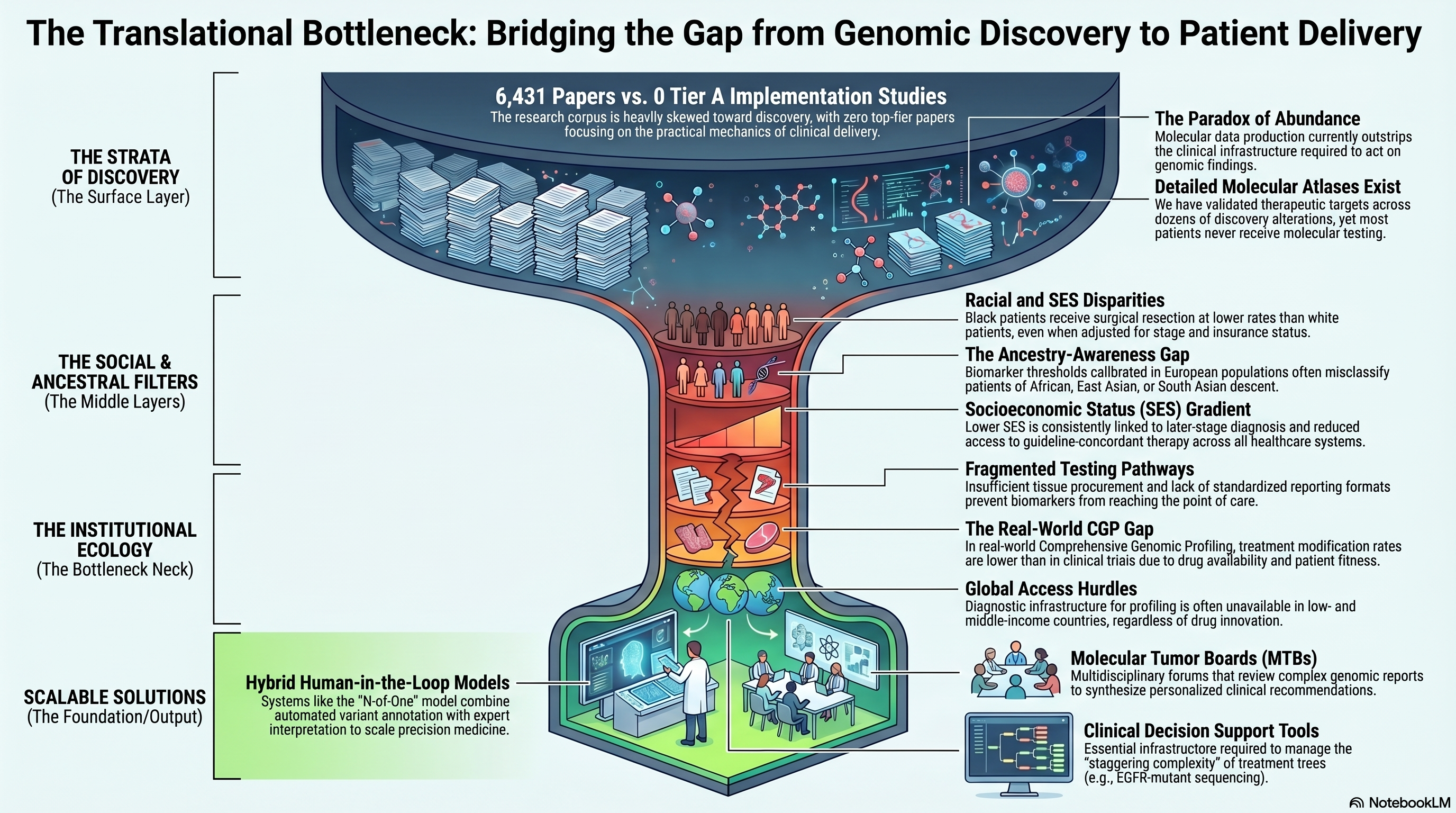 Infographic generated via NotebookLM from the chapter source material.
Infographic generated via NotebookLM from the chapter source material.
Implications for the Manuscript
This theme occupies a unique position in the corpus: it is the only major theme with no Tier A papers, reflecting a systematic undervaluation of translational and equity research relative to discovery science. The manuscript should leverage this bibliometric asymmetry as evidence of a structural gap in the field. Key recommendations include: (1) advocacy for ancestry-aware biomarker development and validation in diverse populations; (2) investment in scalable molecular tumor board infrastructure, including hybrid computational-human interpretation platforms; (3) integration of real-world evidence from comprehensive genomic profiling into treatment decision support systems; and (4) explicit acknowledgment that the translational bottleneck is not a knowledge deficit but an implementation deficit requiring systems-level solutions. The 67 Tier B papers provide sufficient material for a substantive review section, but the manuscript should note the paucity of high-impact translational research as itself diagnostic of the field's priorities.