Artificial Intelligence and Machine Learning in Lung Cancer
"We have entered an era in which algorithms can see what pathologists cannot, predict what clinicians dare not, and integrate what no single human mind could hold — yet the chasm between algorithmic promise and clinical proof remains vast."
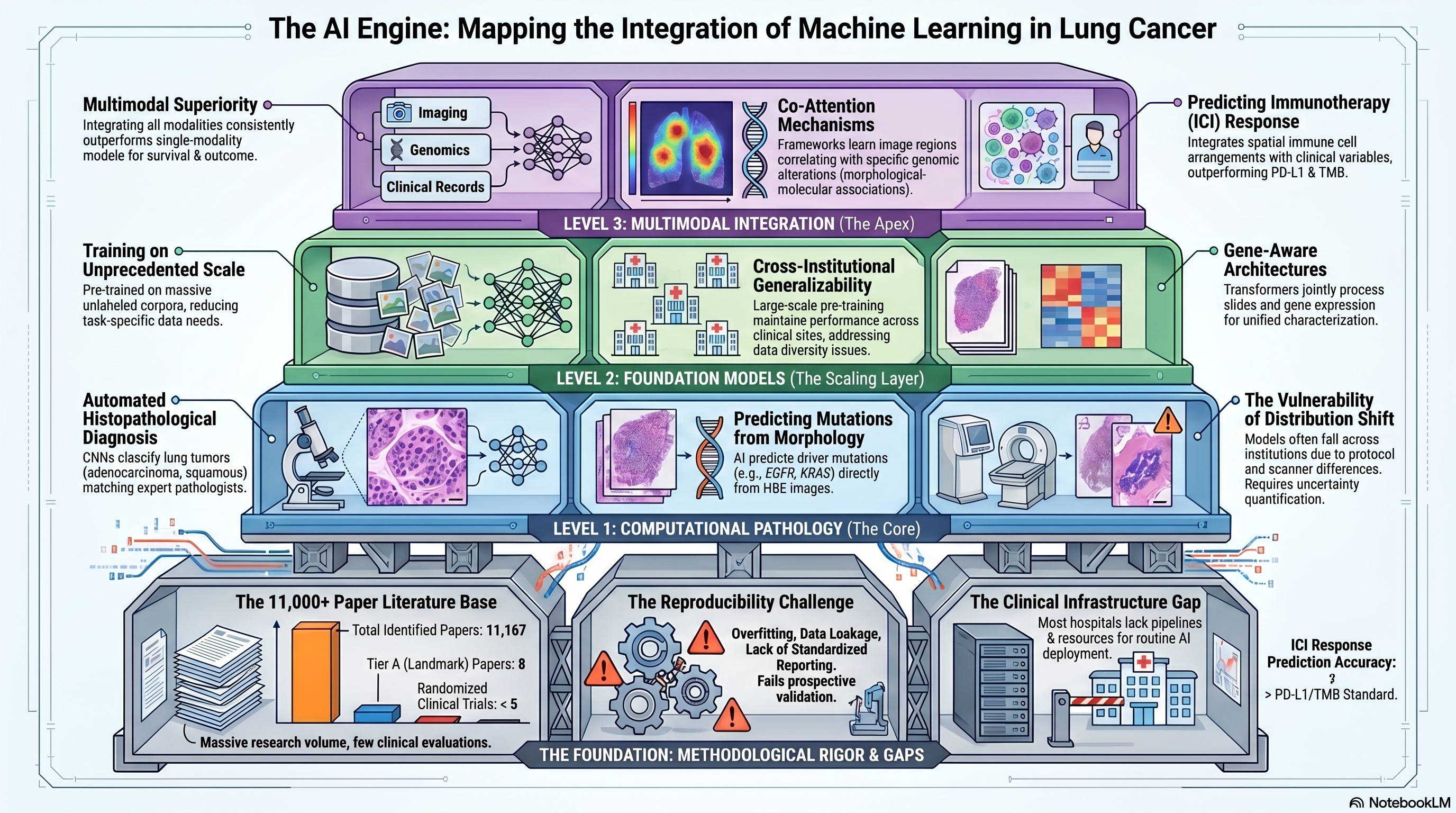 Infographic generated via NotebookLM from the chapter source material.
Infographic generated via NotebookLM from the chapter source material.
Literature base: 11,167 papers identified | 8 Tier A (landmark/highly cited) | 803 Tier B (significant contributions)
Artificial intelligence and machine learning have emerged as transformative technologies in lung cancer research, spanning applications from automated histopathological diagnosis to multimodal outcome prediction. The scale of the literature — over 11,000 papers and climbing rapidly — reflects both the genuine promise of these approaches and the considerable hype that surrounds them. At their best, AI/ML methods extract clinically actionable patterns from complex, high-dimensional data that elude traditional statistical analysis; at their worst, they produce overfit models that perform brilliantly on training data but fail to generalize across institutions, patient populations, and clinical contexts. This chapter examines the major advances in AI/ML for lung cancer, from foundational work in computational pathology through the rise of foundation models and multimodal learning, with particular attention to the methodological rigor — or lack thereof — that underpins published claims.
Computational Pathology: Where Deep Learning First Proved Its Worth
The application of deep learning to histopathology images marked a pivotal moment for AI in oncology. Coudray and colleagues demonstrated that convolutional neural networks trained on whole-slide images of lung cancer could classify tumors as adenocarcinoma, squamous cell carcinoma, or normal tissue with performance approaching that of expert pathologists [PMID: 30224757]. More provocatively, their networks could predict the presence of specific driver mutations — including EGFR, KRAS, and STK11 — directly from hematoxylin and eosin (H&E)-stained tissue images, suggesting that morphological features visible to deep learning but not to the human eye encode information about the underlying molecular state of the tumor. This finding catalyzed a wave of research exploring the relationship between tumor morphology and molecular biology, and established computational pathology as a legitimate bridge between histological and molecular characterization.
The clinical deployment of such models, however, demands a level of robustness and uncertainty quantification that initial studies did not provide. Dolezal and colleagues addressed this gap directly, demonstrating that deep learning models in computational pathology are vulnerable to distribution shift — differences in tissue processing, staining protocols, and scanner characteristics between training and deployment settings — and that without explicit modeling of prediction uncertainty, these models can fail silently, producing confident but incorrect classifications [PMID: 36323656]. Their work introduced frameworks for calibrating model uncertainty and detecting out-of-distribution samples, essential capabilities for any AI system intended for clinical use. Courtiol and colleagues further illustrated the clinical potential of deep learning in thoracic malignancies by developing a model for malignant pleural mesothelioma that predicted patient survival from H&E images, identifying histological features associated with prognosis that were not captured by conventional pathological grading systems [PMID: 31591589]. Their approach of using attention mechanisms to highlight the most prognostically informative tissue regions provided a degree of interpretability that is critical for clinical acceptance.
Foundation Models: Scale and Generalization
The emergence of foundation models — large neural networks pre-trained on massive unlabeled datasets and then fine-tuned for specific tasks — has introduced a paradigm shift in computational pathology and medical image analysis. Wang and colleagues developed a pathology foundation model trained on an unprecedented corpus of whole-slide images spanning multiple cancer types and tissue preparations [PMID: 39232164]. This model achieved state-of-the-art performance across a diverse range of downstream tasks, including tumor classification, mutation prediction, and survival estimation, with substantially less task-specific training data than traditional supervised approaches. The foundation model paradigm addresses one of the central challenges in medical AI — the scarcity of labeled training data — by learning general-purpose visual representations from unlabeled images that can be transferred to specific clinical tasks with minimal additional annotation.
Yang and colleagues pushed this concept further with a generalizable foundation model designed specifically for medical image analysis, demonstrating robust performance across multiple cancer types, imaging modalities, and clinical sites [PMID: 40064883]. Their model was explicitly evaluated for cross-institutional generalizability, addressing the distribution shift concerns raised by Dolezal and colleagues, and showed that large-scale pre-training significantly mitigates the performance degradation typically observed when models are applied to data from institutions not represented in the training set. For lung cancer specifically, foundation models offer the tantalizing possibility of developing AI tools that generalize across the enormous diversity of tissue preparation and imaging protocols encountered in real-world clinical practice.
The integration of molecular and imaging data through gene-aware architectures represents another frontier. Wang and colleagues introduced Gene Swin, a model that combines the Swin Transformer architecture for processing whole-slide images with gene expression data to jointly predict molecular subtypes and clinical outcomes [PMID: 40515391]. By explicitly modeling the relationship between histological features and gene expression patterns, Gene Swin bridges the gap between computational pathology and molecular profiling, offering a multimodal approach that leverages complementary information from both data types. This architecture is particularly relevant for lung cancer, where the integration of histological and molecular data is essential for comprehensive tumor characterization but is rarely achieved in a unified analytical framework.
Multimodal AI: Integrating Across Data Types
The natural extension of single-modality AI models is multimodal learning, which integrates data from multiple sources — imaging, genomics, clinical records — to produce more comprehensive predictions. Chen and colleagues developed a multimodal framework that combines histopathology images with genomic features using a co-attention mechanism, allowing the model to learn which image regions are most informative in the context of specific genomic alterations and vice versa [PMID: 35944502]. Applied to multiple cancer types including lung cancer, their model demonstrated that multimodal integration consistently outperformed either modality alone for survival prediction, and the attention maps provided interpretable visualizations of the morphological-molecular associations learned by the network. This work established multimodal AI as a viable strategy for integrating the disparate data types that characterize modern oncology and laid the groundwork for more sophisticated architectures that incorporate additional modalities.
Arshad and colleagues provided a comprehensive review of AI applications across the lung cancer continuum, from screening and early detection through diagnosis, treatment selection, and outcomes prediction [PMID: 41463234]. Their synthesis highlighted the remarkable breadth of AI applications in lung cancer — encompassing radiology, pathology, genomics, and clinical decision support — while noting that the vast majority of published models have not been validated in prospective clinical settings. They identified a striking asymmetry in the literature: thousands of studies report novel AI models for lung cancer, but fewer than a handful have been evaluated in randomized clinical trials, and even fewer have been integrated into routine clinical workflows. This gap between algorithmic development and clinical implementation represents the central challenge for the field.
Predicting Treatment Response: Immunotherapy and Beyond
The prediction of response to immune checkpoint inhibitors (ICI) has emerged as one of the highest-value applications of AI in lung cancer, driven by the clinical urgency of identifying patients who will benefit from these expensive and potentially toxic therapies. Rakaee and colleagues developed an AI-based predictor of ICI response that integrates histopathological features with clinical variables, demonstrating superior discrimination compared to PD-L1 immunohistochemistry and tumor mutational burden, the two most widely used clinical biomarkers [PMID: 39724105]. Their model identified morphological features of the tumor microenvironment — including the spatial arrangement of immune cells relative to tumor cells — that were associated with treatment benefit, providing both predictive utility and biological insight. The clinical importance of this work cannot be overstated: current biomarkers for ICI selection have positive predictive values of only 30-40%, meaning that the majority of patients selected for treatment based on PD-L1 or TMB do not derive meaningful benefit.
The multicenter validation of AI models is essential for establishing their clinical credibility, yet remains the exception rather than the rule. Thiesen and colleagues conducted a rigorously designed multicenter evaluation of an AI model for lung cancer, training on data from multiple institutions and testing on held-out sites to assess generalizability under realistic conditions [PMID: 41481196]. Their study revealed that while the model maintained good overall performance across sites, there was meaningful variation in accuracy that correlated with differences in patient demographics, tissue processing protocols, and imaging equipment. This finding underscores the importance of diverse, multi-institutional training data and of reporting performance stratified by clinical site rather than relying on aggregate metrics that can obscure site-specific failures.
Standards, Reporting, and the Reproducibility Challenge
The rapid growth of the AI/ML literature in lung cancer has outpaced the development of reporting standards and quality benchmarks, creating a landscape in which the methodological rigor of published studies varies enormously. Bakas and colleagues, working through the AI-RANO initiative, developed standardized guidelines for the evaluation of AI models in neuro-oncology that have broader applicability across cancer types [PMID: 39481415]. Their framework specifies requirements for dataset documentation, model transparency, statistical evaluation, and clinical relevance assessment that address many of the methodological shortcomings prevalent in the AI oncology literature. The adoption of such standards in the lung cancer AI community would substantially improve the reproducibility and credibility of published work.
The reproducibility challenge in AI/ML for lung cancer is multifaceted. Beyond the well-known issues of overfitting and data leakage, there are subtler problems related to the handling of confounders (for instance, models that learn to predict outcomes from artifacts of tissue processing rather than genuine biological features), the absence of prospective validation, and the use of inappropriate performance metrics. Many published models report area under the receiver operating characteristic curve (AUROC) as the primary performance metric, despite the fact that AUROC can be misleading in the setting of class imbalance — a situation that is common in clinical oncology, where events such as treatment response or recurrence may occur in a minority of patients.
Where Consensus Exists and Where the Field Disagrees
There is broad consensus that AI/ML has demonstrated proof-of-concept value in lung cancer across multiple applications, from histopathological classification to outcome prediction. The superiority of deep learning over traditional machine learning approaches for image analysis tasks is generally accepted. The importance of external validation, uncertainty quantification, and interpretability for clinical translation is acknowledged in principle, if not always in practice.
Substantial disagreements persist, however. The clinical readiness of AI models for lung cancer remains fiercely debated, with optimists pointing to the strong performance of foundation models and multimodal approaches and skeptics emphasizing the near-complete absence of prospective validation. The question of whether AI should augment or replace human decision-making in oncology is philosophically contentious: some investigators envision AI as a clinical decision support tool that informs but does not supplant clinical judgment, while others argue that the full benefit of AI will only be realized when algorithms are given greater autonomy in clinical decision-making. The appropriate regulatory pathway for AI-based diagnostic and prognostic tools in oncology is also unsettled, with existing frameworks designed for static medical devices struggling to accommodate the dynamic, continuously learning nature of modern AI systems.
Critical Gaps
The most critical gap in the field is the near-total absence of prospective, randomized evidence that AI tools improve clinical outcomes in lung cancer. While retrospective studies have demonstrated impressive discriminative performance, the translation to prospective clinical benefit requires demonstration that AI-guided decisions lead to better outcomes than standard-of-care approaches — a bar that has not yet been met for any AI application in lung cancer. Second, the equity implications of AI in lung cancer are underexplored: models trained predominantly on data from academic medical centers serving predominantly White patient populations may perform poorly — and could even exacerbate disparities — when deployed in more diverse clinical settings. Third, the integration of AI with multi-omics data is still in its early stages, with most published models relying on single data modalities; the development of truly multimodal AI systems that seamlessly combine imaging, genomic, proteomic, and clinical data remains an aspiration rather than a reality for most research groups. Fourth, the interpretability of complex AI models — while improving — remains insufficient for many clinicians, who are understandably reluctant to act on predictions from opaque algorithms. Finally, the technical infrastructure required to deploy AI models in clinical settings — including data pipelines, computational resources, and integration with electronic health records — is lacking at most hospitals and cancer centers.
Implications for the Manuscript
This chapter completes the thematic arc of Part 2 by showing how AI/ML provides the computational engine needed to operationalize the molecular heterogeneity and multi-omics integration discussed in preceding chapters. The progression from computational pathology through foundation models to multimodal AI mirrors the increasing ambition of the field, from automating existing tasks (histological classification) to enabling entirely new capabilities (predicting molecular features from morphology, integrating imaging with genomics). The critical gaps identified here — particularly the need for prospective validation, equitable model development, and clinical infrastructure — will be central to the review's recommendations. The discussion of foundation models and multimodal AI will inform the manuscript's vision for the future of precision oncology in lung cancer, where AI serves as the integrative layer that connects disparate data types into coherent, actionable patient profiles.